Research Projects
Advancing peptide therapeutics through innovative design and rigorous scientific methodology
Current Research Focus
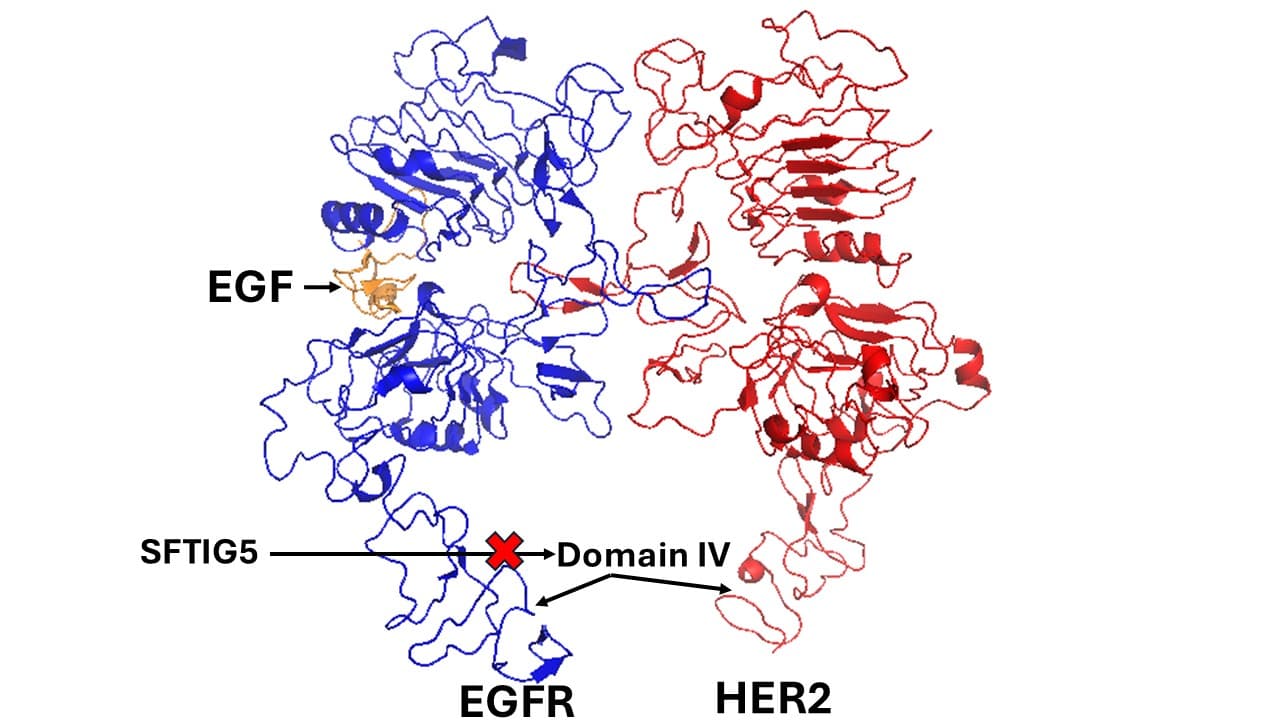
Our research focuses on developing orally bioavailable grafted peptides targeting EGFR receptor dimers for non-small cell lung cancer therapy. We integrate computational modeling (PyMOL, MOE, Discovery Studio, NAMD, GROMACS) with experimental validation using SPR, HPLC, kinase assays, and confocal imaging, complemented by in vivo studies in xenograft and PDX models.
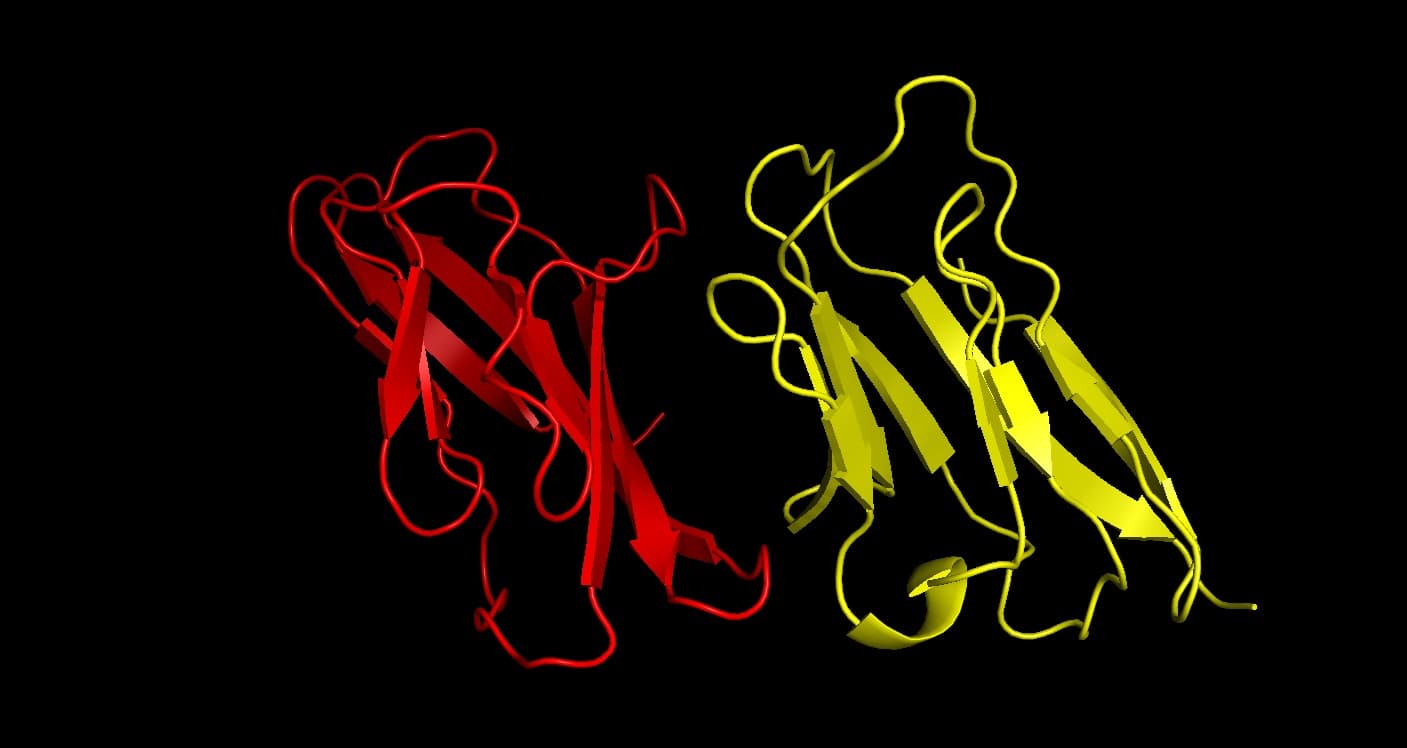
Our research involves soluble peptide and peptidomimetic inhibitor for inhibition of Costimulatory CD2-CD58 nexus thus preventing against autoimmune disease like Rheumatoid Arthritis. We design peptide-based therapeutics that can modulate protein-protein interactions critical to immune system regulation.
Collaborative Research
Contributed to the development and testing of the NIGMS Structural Biology and Drug Discovery Sandbox Model, applying computational pipelines to evaluate molecular interactions and therapeutic targets. This work advances the integration of computational methods in drug discovery workflows.
Methodological Expertise
- Molecular Dynamics: GROMACS, AMBER, CHARMM force fields
- Docking & Scoring: AutoDock Vina, Glide, FlexX
- Free Energy: FEP, TI, Umbrella sampling
- QSAR/QSPR: Machine learning models for activity prediction
- RNA-seq Processing: Cell Ranger on HPC clusters
- Differential Expression: DESeq2, EdgeR in R
- Pathway Analysis: GSEA, IPA
- Visualization: Loupe Browser, Space Ranger
- Statistical Analysis: GraphPad Prism, MATLAB, JMP
- SPPS: Fmoc/tBu strategy, microwave-assisted synthesis
- Cyclization: Disulfide, lactam, click chemistry
- Stapling: Hydrocarbon, triazole, thioether bridges
- Modifications: Unnatural amino acids, PEGylation
- Binding Kinetics: SPR (Biacore), ITC, MST
- Structural Analysis: CD, NMR, X-ray crystallography
- Stability Studies: Proteolytic, thermal, chemical
- Cell Studies: Uptake, localization, cytotoxicity
